This protocol was designed for a Brain Canada project acquiring MRI data at the MNI, but with the intention to make it freely available so that other groups could benefit from our efforts, and so that ultimately the various groups could share data. It is already being used by groups in Canada, Europe, and the US.
The protocol design is intended to maximize success in collecting data in populations in which movement could be a problem, e.g. children, the elderly, or individuals with disorders. To this end, the sequences are kept as short as possible, both to minimize the chance of movement during a scan, and also to allow a sequence to be repeated, if corrupted by movement. This aspect of the protocol relies, in part, on the acceleration capabilites of the Siemens Prisma 3T.
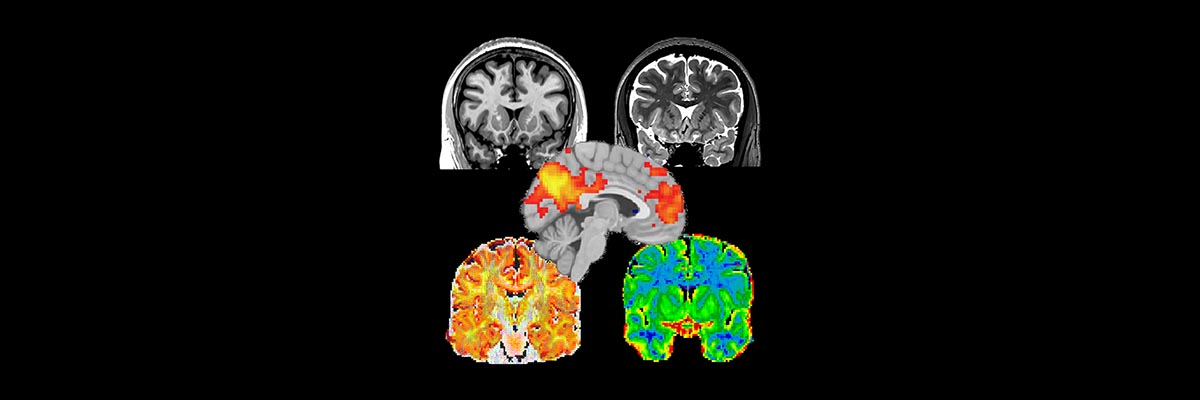
The protocol generates T1-weighted data; T2-weighted data; diffusion data suitable for both DTI and NODDI analyses, as well as for probabilistic tractography, and the corresponding distortion correction data; MT-sat data and a B1-map that together with the NODDI data can be used to produce a g-ratio map; and resting state fMRI data. A description of the sequences that produce each sort of data, with notes for the MR technicians, can be found here.
The protocol parameters can be found here; the loadable exar files for a Siemens Prisma 3T can be found here and here; and the diffusion vectors can be found here. The protocol uses strictly Siemens sequences (no C2Ps are required). The design is intended for use with a 64-cannel head coil (though portions of it have also been tested with a 32-channel head coil).
The protocol was designed by John Lewis with help from Ilana Leppert, Michael Ferreira, and the MRI technicians at the MNI’s Brain Imaging Center. Questions should be directed to Dr. Lewis